I’ve copied the following questions from private email (with permission) so that they can be answered publicly.
On 5/17/20 11:49 AM, Johnny Ghibril wrote:
> Good morning David,
> Thank you for getting back to me so quickly and sorry for the delayed
> response. You gave us quite a bit to look into and I wanted to make sure
> we are well educated before taking any of your, or the community's,
> time. We will join the conference call this week, but since there's a
> set agenda, we also don't want to sidetrack the group so I've outlined
> some basic questions below.
Our weekly teleconference has been temporarily interrupted by COVID-19 and other activities, but we do plan to return to them as soon as we can. If you are not already on the HL7 ITS list or the W3C Healthcare and Life Sciences list then I recommend subscribing, because that’s where we announce our teleconferences:
https://lists.w3.org/Archives/Public/public-semweb-lifesci/
> I am not sure where the best forum is to get
> them answered and would appreciate your guidance:
>
> 1. The RDF format is designed using properties like fhir:nodeRole,
> fhir:treeRoot
Those two were needed because RDF, being graph oriented rather than tree oriented, does not have any notion of a root/top/starting element. Therefore, when converting FHIR JSON or FHIR XML to FHIR RDF, we needed a way to indicate what the root element was. This need stems from the overall requirement that conversion of FHIR between JSON, XML and/or RDF must be fully round-trippable – no information gained or lost in the process. It is conceivable that there could be some other way around this problem, but the most straightforward solution seemed to be to introduce fhir:nodeRole fhir:treeRoot.
> and fhir:index. We are not sure why such specific
> representations were used, why not a “standard” rdf/owl
> representation with uri references?
fhir:index is used to indicate the ordering of items in a list, which in FHIR is considered semantically significant (unless explicitly stated otherwise). Since RDF is an unordered set of triples, something explicit is needed in RDF to indicate the ordering of items in a list. We could have used an rdf:List for these items, but they are notoriously unfriendly to SPARQL queries. There are several other proposed list representations available in the RDF/OWL community, but alas, none of them are standard. So we chose fhir:index, in the hope that someday a standard mechanism will be adopted in the RDF/OWL community, and we could later switch to that.
With that said, we are currently thinking about the next version of FHIR RDF (which we’re calling R5), and this is one of the issues that we are reconsidering. Any input would be welcomed.
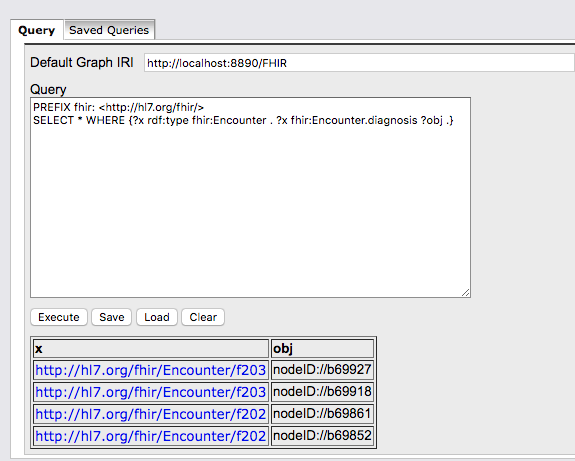
> 2. The ttl format contains anonymous/blank nodes which cannot be
> handled by a triplestore or a tool like Protegé. We did ask this
> question in the FHIR community forum jere:
> http://community.fhir.org/t/anonymous-blank-nodes-in-ttl-files-vs-resource-nodes/1940
> But unfortunately not much of a response there. These blank nodes
> are a problem because they seem to corrupt the representation and
> “co-reference” relations, and so using SPARQL queries seem to be
> compromised. It seems therefore we have to write some code to map
> those nodes to proper URIs. Is that correct? Have you or the
> community already solved this?
Great question. Those bnodes were used because FHIR allows (nearly) EVERYTHING to be extended – including primitive types like boolean. So the bnodes are there to enable extensions to be inserted. HOWEVER, as you have noted – and Harold Solbrig has also noted – the downside of that decision is that it makes SPARQL queries more cumbersome in the most common cases. Consequently, this is another design decision that we are reconsidering for the next version of FHIR RDF, and your input would be welcomed.
> 3. We are investigating using FHIR-RDF to be able to use semantic web
> standards. However it seems the HL7/FHIR has not seen much traction
> there. Any thoughts on this? Is this intentional or simply a
> yet-to-be-explored avenue?
Interestingly, we have observed that there is minimal interest in FHIR RDF from within the FHIR community, but there is interest from outside the FHIR community, typically by researchers who want to be able to combine EHR data (via FHIR) with other biomedical data that they have. The general FHIR community is MUCH larger though, and that means that we have almost no influence on the design of FHIR: we just have to accept it as we get it, and focus on the problem of representing it in RDF. However, an exception can happen if we discover a design issue and convince the general FHIR community that it will also affect them.
> 4. As I mentioned to Harold in my email, we are a young startup and
> starting to explore the space. Any recommendations you have for us
> are welcome. For example, do you see FHIR-RDF a
> recommended/promising route in your opinion? Or is the “standard” to
> programmatically handle FHIR representations via an API such as HAPI?
I don’t think I could make a recommendation one way or another without knowing more about your specific circumstances, but as a general rule I’d say it mainly depends on whether you are already using RDF for other reasons. If so, it probably makes sense. If not, probably not.
> We would be happy to get on an introductory call if that is more
> efficient/appropriate.
We would love to have you join a call and tell us more about what you are doing, and have your participation in improving the next version of FHIR RDF. I don’t yet know when our next teleconference will be, but if you subscribe to the list I mentioned above, you’ll get the notification.